Applications: - Peptide Sequencing
In the last 15 years
or so, ESI-CID-MS/MS analysis
has seen increasingly wide application in the area of proteomics -
especially peptide sequencing and protein identification. CID
fragmentation of a singly or doubly protonated ion ( [M+Hn]n+
- where n=1 or 2) of a peptide will yield a series of fragment ions
resulting from the cleavage of the peptide at any of the C-C bonds
between amino-acid residues
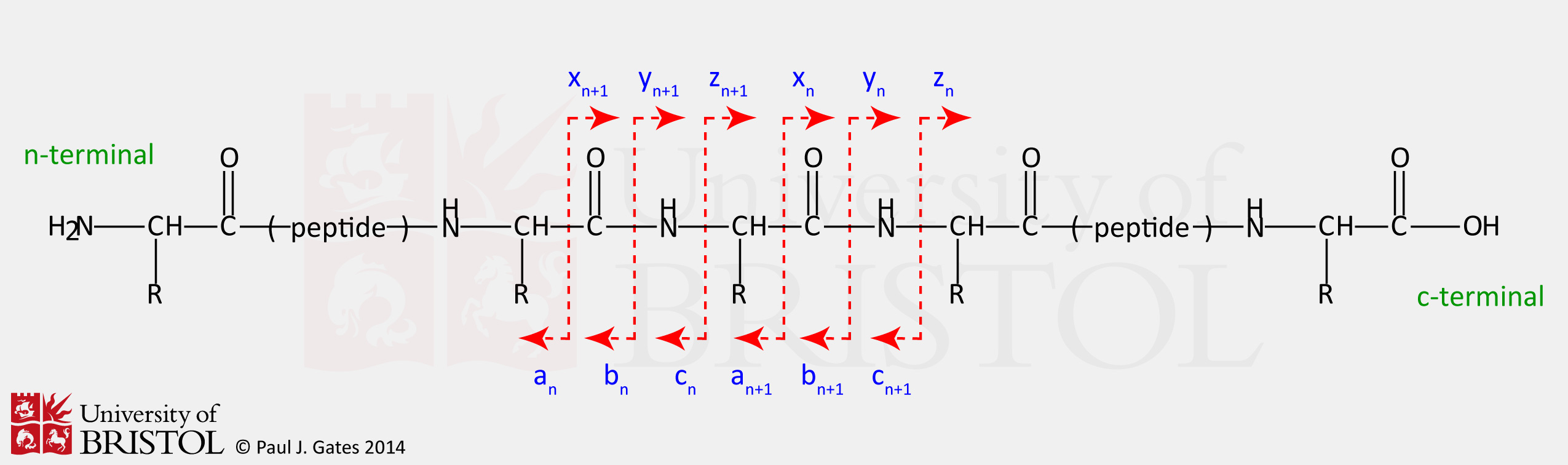
(see figure 1).
In CID, 'b' and 'y' ions are most often observed along with 'a' ions resulting from the sub-fragmentationof 'b' ions. The other possible types of sequence ions can be produce from other fragmentation methods. The mechanism of peptide fragmentation involves one of the proton charges, normally resident on an amide, being mobilised onto any one of the peptide amide bonds (see figure 2). This subsequently weakens the amide bond resulting in cleavage. Assuming that a heterogeneous population of protons exists in the precursor ion, this cleavage results in a complementary series of 'b' and 'y' ions. It is also possible to form internal immonium ions. Certain residues can enhance the fragmentation at their position, especially those that contain an extra labile proton on their side group that can act as an additional mobile proton. Similar mechanisms can be drawn for the formation of all the various types of sequence ion.

Figure 2: Scheme showing peptide fragmentation initiated via the mobile proton mechanism to form a 'y' type ion.

Figure 1: Nomenclature and bond
cleavage sites for all the various types of peptide fragment
(sequence) ions.
In CID, 'b' and 'y' ions are most often observed along with 'a' ions resulting from the sub-fragmentationof 'b' ions. The other possible types of sequence ions can be produce from other fragmentation methods. The mechanism of peptide fragmentation involves one of the proton charges, normally resident on an amide, being mobilised onto any one of the peptide amide bonds (see figure 2). This subsequently weakens the amide bond resulting in cleavage. Assuming that a heterogeneous population of protons exists in the precursor ion, this cleavage results in a complementary series of 'b' and 'y' ions. It is also possible to form internal immonium ions. Certain residues can enhance the fragmentation at their position, especially those that contain an extra labile proton on their side group that can act as an additional mobile proton. Similar mechanisms can be drawn for the formation of all the various types of sequence ion.

Figure 2: Scheme showing peptide fragmentation initiated via the mobile proton mechanism to form a 'y' type ion.
Each amino acid residue gives a
characteristic mass loss (the residue mass) enabling the
sequencing of the peptide. This
is usually performed automatically by software sequencing
algorithms.Large proteins can by digested into smaller
peptide/protein fragments
by use of a suitable enzyme, and HPLC-MS/MS methods employed to
perform the sequencing. Databases of protein sequences are so vast
that complete sequencing of a protein is rarely required. Partial
sequencing will usually give the 'correct' answer at a high degree
of
certainty. It is not even always necessary to resolve ambiguous
sequence information (for example the isomeric Ile and Leu or the
isobaric Lys
and Gln).