|
The Manganese-calcium oxide cluster of Photosystem II and its assimilation by the Cyanobacteria James D. Johnson M.S.
Key: Hyperlink, Pop-up window Introduction:
This month's MOTM is an inorganic cubane complex known as the manganese-calcium oxide
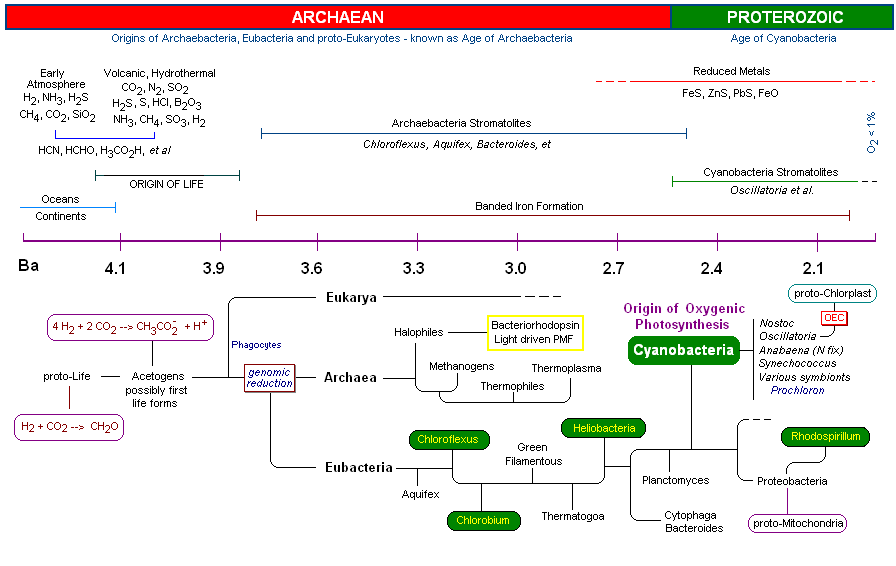
cluster, commonly referred There are two photosystems, PSII and PSI, which communicate with one another via an electron transport chain known as the "Z scheme" of photosynthesis. Electrons are initially extracted from water, one at a time (1 photo per electron) at the site of the OEC, and finally released to a soluble ferredoxin complex after passing through PSI (where it is further re-energized by a second photon). Ferredoxin then reduces Fd-NADP oxidoreductase, which reduces oxidized NADP, which then reduces 1,3-bisphosphoglycerate. This reduction step regenerates Ribulose-5-phosphate followed by incorporation of CO2 by Ribulose-bisphosphate carboxylase (Rubisco) - the most abundant protein on the planet). The reduction of CO2(carbon fixation) is known as the "dark reaction" or Calvin Cycle (sugar production). In algae and plants the thylakoid membranes are found partially stacked (grana) where PSII occurs in higher concentration than PSI (the stacking of the thylakoids may be related to an increase in the photon transfer efficiency by the surrounding light harvesting chlorophyll complexes - in the pop up of the light harvesting complexes, notice the concentric arrangement of protein, chlorophyll (and carotenoids), centered around the PSII reaction center. Recently the three dimensional atomic structure of the manganese-calcium oxide complex was determined by X-Ray diffraction. (3) The manganese cluster is essentially an inorganic cubane (cubical) structure found in inorganic crystals made of ranciéite or hollandite [Mn4CaO9*3H2O]. These minerals are believed to have been assimilated during the early Archaean period by the Cyanobacteria approximately 3,200 - 2,800 million years ago (or 3.2 - 2.8 Ga=giga years ago). The assimilation of the manganese-calcium oxide complex by the Cyanobacteria is one of several examples of metallo-catalysts incorporated by living systems. Two other inorganic metallo complexes assimilated by early prokaryotes (and are now ubiquitous redox centers in living systems) include the cubic iron-sulfur centers [Fe4-S4] [image] (originating as greigite (Fe4S4][SFeS]2)(4), such as ferredoxin, and the nickel-iron centers [NiFe][image], such as Hydrogenases. The successful assimilation of the OEC by early Cyanobacteria resulted in an explosion of this group which became the predominant life form for 2 billion years. Unlike other photosynthetic bacteria (discussed briefly below) the Cyanobacteria are "oxygenic photosynthesizers" capable of oxidizing water as a substitute for other electron donors such as H2S and organics (acetate, succinate, e.g.), producing molecular oxygen as a by-product. Oxygen irreversibly changed the course of life on earth by releasing O2 in great quantities (changing, for example, the composition of the atmosphere as well as earth's geology). During this time the Cyanobacteria successfully dispersed globally and diversified through evolutionary speciation. (5) Finally, member(s) of the Cyanobacteria symbiotically merged with eukaryotes at about 2.0 Ga to become the chloroplast of all algae and higher plant groups on the earth today. The arrival of the OEC of PSII resulted in the most profound impact on the history of evolution. Indeed, the OEC is single-handedly responsible the rise of all plants and animals on the earth today. 1.
First Life - the extremophilic Archaebacteria (4.0 - 3.5 Ga)
1. First Life - The Extremophilic Archaebacteria (4.0 - 3.5 Ga) Early Earth:
It is estimated that earth's oceans, continents and atmosphere required approximately 600 million years to stabilize (4.6 - 4.0 Ga) to
the point where liquid water accumulation and continental crust accretion were possible. (3)
Some reports put this time as early as 4.4 Ga. (4).
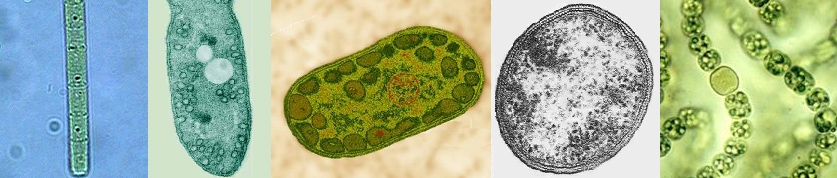
Early chaotic surface events, though becoming less frequent with time, included meteor bombardment, unstable temperatures, strong lunar forces, tides and storms, risings oceans, reduced sunlight intensity, short day length, UV, X-Ray and other high energy irradiation, volcanic and geologic upheavals and an ever changing atmospheric chemistry (6). Earth's atmosphere at the beginning of the Hadean period (4.6 - 4.2 Ga) was strictly reducing (lacked oxygen) (7), having measurable levels of H2, NH3, CH4 and H 2S. Outgassing by volcanic and thermal vent activity released reduced metals, especially Iron, as well as Phosphorous, Sulfur, H2, H2S, CH4 and CO2 resulting in an acidic ocean. (8) Due to occasional extreme temperature conditions, exceeding 800oC (9), as well as active solar flaring (10) and other high-energy events, it is believed that CH4 and NH 3 would have been heat destroyed. So long as molecular oxygen was absent, the earth's atmosphere remained in a reduced state. (11) By around 4.2 Ga, the earth's atmosphere was becoming relatively stable and consisted primarily of volcanic gas. At this time the atmosphere was primarily water vapor (94%), CO (4%), N 2(4%), CO, and SO 2 (12). Fig 2. represents an image of the earth at about 4.0 Ga, when oceans were deposited, continents formed, and periodic meteorite strikes were diminishing. Fig 3 is an artist's rendition of the earth at about 3.8 Ga (note that the moon is much closer) which corresponds to the time that Life first appeared on the earth. (13) Hydrothermal vents and volcanic outgassing created a large reservoir of oceanic Fe(II) and FeS. Oxidation of metals by UV radiation in oceanic surface waters may have resulted in an accumulation of Fe and Mn oxide precipitates, which subsequently settled out. (14) These early oxidized metals, such as Fe(III), may have served as life's first electron acceptors. First Life: The first organisms were extremophilic chemotropic prokaryotes (that is, tolerated extreme environmental conditions such as high heat, high salt concentration, high and low pH, etc). The extremophiles are now known to have been early members of the domain Archaebacteria, although it is argued that simultaneous evolution of the eubacteria as well as the eukaryotes paralleled these early extremophiles (15). Many species of the archaebacteria survive today (such as the acetogens, methanogens, thermophiles and halophiles). These organisms, like true bacteria, lack a nucleus, as well as organized internal membrane structures, vacuoles and internal organelles . The early archaebacteria did however have an amazing array of biochemical pathways to derive energy, acquire carbon, nitrogen, phosphorous, sulfur, hydrogen, bicarbonate and various organics as well as the requisite minerals needed for growth. In addition they had RNA and DNA, giving them the ability to store and pass on genetic information necessary to build enzymes to carry out the multitude of necessary biochemical processes for life. In addition, the early prokaryotes had the capacity (as extant members of the Archaebacteria have today) to "detect" and "respond" to changes in their environment, such as redox potentials, nutrient levels, physical events, light and temperature variances, et al. The ability to respond to changes in their immediate environments allowed them to adjust, tolerate, thrive, reproduce and evolve. The various hypothetical pathways which led to earth's first life forms from proto-life is an area of active and intense research. It is believed that the first organisms were acetogens or closely related methanogens [see Timeline below]. The oldest fossils of life on earth are found in microfossils (3.8 Ga) and stromatolites (3.5 Ga). As shown in (Fig 5, 6 and 7) Stromatolites appeared in the early shallow waters of the earth approximately 3.8 Ga. The artist's rendition in Fig. 5 (below, left) of the earth at that time accurately depicts the volcanic outgassing and undersea hydrothermal vent activity which would have been prevalent near the end of the Hadean eon. The earliest stromatolites were probably pre-Photosynthetic eubacteria such as Chloroflex and Chlorobium, both photosynthetic archaean anaerobic thermophiles which preceded the Cyanobacteria.
The assimilation of the OEC by the cyanobacteria laid the ground work for the Cambrian explosion, and for the evolution of the animal and plant kingdoms. Cyanobacteria, by assimilating the OEC and releasing molecular oxygen, changed the geological landscape of the earth, as well as its atmosphere and oceans. All other organism would respond in abeyance, either taking up oxygenic respiratory or related metabolism, or recess into niches devoid of oxygen's diffusional persistence. Recent genomic mapping and macromolecular resolution (especially by X-Ray diffraction) as well as a host of other approaches (kinetic studies, et al.) has led to an explosion of research on the evolution of life. The recent results of these efforts, brought about by the computer and advanced technological research capabilities, have been staggering. Previous concepts of taxonomy, based primarily on morphology and physiology, have been supplanted by the work of Woese and others who revolutionized the concept of DNA as molecular fossils (16). Woese and his group are responsible for the discovery of the domain Archaea, and in the process laid the ground work for future research in this area (Woese's work gives new meaning to the saying "..when one door is closed, many more are open."). 2. Bacterial Photosynthesis (3.8 - 2.7 Ga) Introduction to the Photosynthetic bacteria: Thirty-six bacterial lineages (Eubacteria) have been identified, of which only five are capable of using chlorophyll-based energy conversion to create a Protonmotive Force (PMF) to drive ATP synthesis and reduce CO2 to sugars. Of the five bacterial groups capable of photoautotrophic photosynthetic growth, four, (the exception being the Cyanobacteria), perform photosynthesis under anaerobic conditions and do not oxidized water to molecular oxygen via the OEC (anoxygenic photosynthesis). Indeed, the Cyanobacteria are the only group of bacteria which have assimilated the OEC necessary for splitting water. Members of the five photosynthetic bacterial families are shown below (these are just example species from each group):
One of the most fascinating areas of biochemistry today is research designed to determine the various evolutionary relationships among these lineages and their evolutionary origins (as well as all of the Eubacteria). Recent findings have revealed that these groups are highly heterophyletic. Scientists now believe that the transfer of genetic information between divergent organisms has been prolific, especially early in evolution, thereby complicating further the genomic mapping of their ancestry (genes may be transferred between divergent organisms though such processes such as conjugation. (17) These events have literally turned the "tree of life" into a three dimensional "web of life." With species dying out in the interim, and evolution spanning more then 3,000 million years, biochemical researchers certainly have their work cut out in the foreseeable future.
All known microorganisms use two functional principles (both mutually exclusive and represent two independent evolutionary developments) for the conversion of light into chemical energy. Chlorophyll-based systems are widespread among members of the domain bacteria and consist of a light-harvesting antenna and reaction centers. In the latter, excitation energy is converted into a redox gradient across the membrane Bacterial reaction center and light harvesting chlorophylls: [LHC section to be completed today, 6-1]
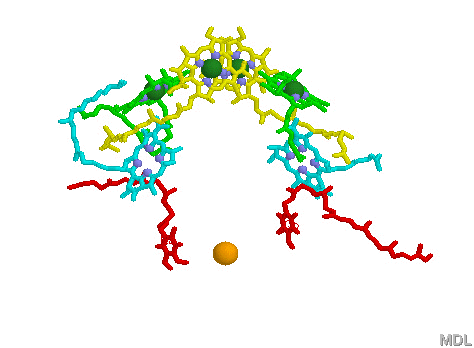
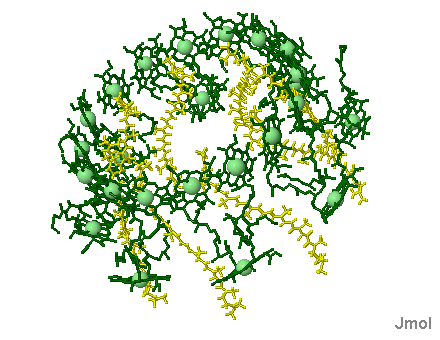
Bacterial reaction center shown above, left. Note the Bacteriachlorophyll reaction center "dimer", a special pair of chlorophyll molecules designed to absorb incoming photons from surrounding light harvesting chlorophyll complexes, shown on the right. Blue chlorophylls shown in the reaction center at left are pheophytins, bacteriachlorophyll minus the Mg atom inside the chlorophyll heme. Red molecules are quinones (the extended tails of the quinones allows for lateral motion and membrane solubility). The electron path begins with the oxidation of the dimer pair of chlorophylls (the reaction center proper) followed by electron transfer through the pheophytins and onto the quinones. The reaction center dimer is then reduced by an external electron sources such as Cytochrome.
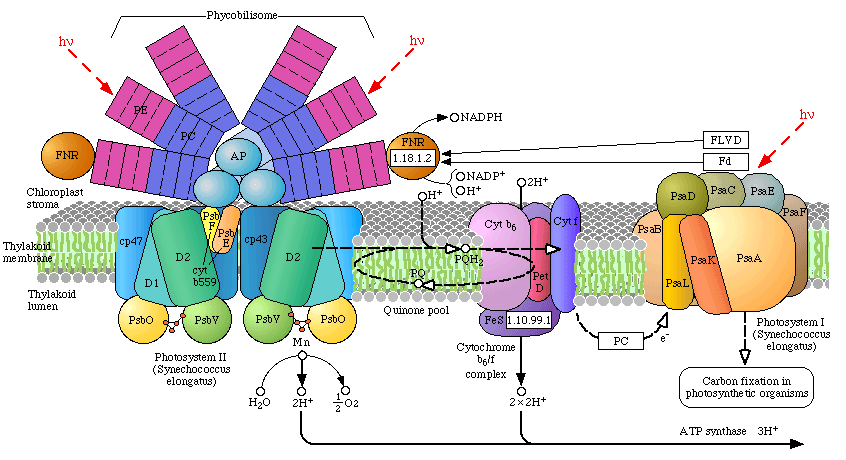
Phycobilisomes: Phycobilisomes are light harvesting pigments found throughout the Cyanobacteria (and in red algae). All other photosynthetic bacteria use the light harvesting complex as described above. Phycobilisomes, which specialize in absorbing deep penetrating green light (more than 1 meter deep, e.g., in marine environments), are capable of re-emitting light in regions which the other photosynthetic pigments can absorb. This would be a very advantageous adaptation in marine environments in which protection from UV radiation might be needed. There is a central core of allophycocyanin (AP) which sits above the photosynthetic reaction center. There are phycocyanin and phycoerythrin subunits which radiate out from this center like thin tubes. The fluorescent pigments which are present in the phycobilisome, such as phycocyanobilin (PC) and phycoerythrobilin (PE) re-emit the green light in regions.
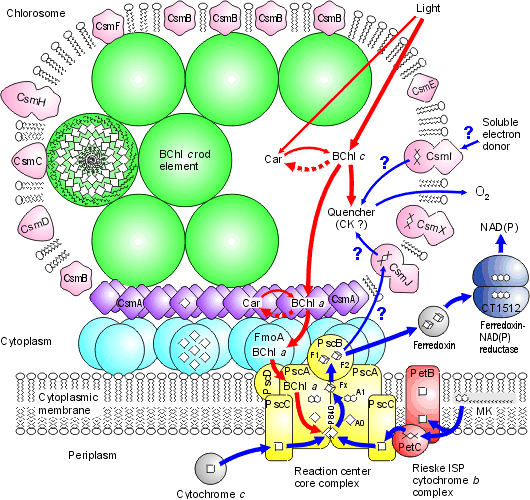
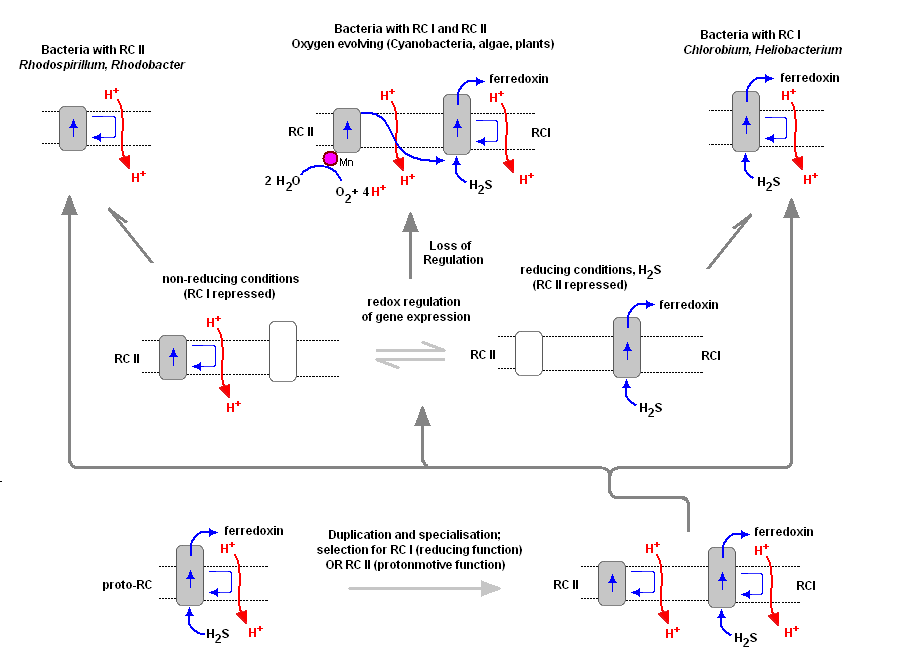
Chlorosomes: Model of the photosynthetic apparatus in Chlorobium tepidum showing "chlorosome" body compiled by Dr. Donald Bryant and associates at Penn State University (Chlorosomes found in the green sulfur bacteria, family Chlorobiaceae). Light and excitation energy transfer is shown in red, electron transfer is shown in blue. Proteins and chlorosome rod elements are shown in color and some cofactors are indicated as white symbols. Diamonds represent individual BChl c molecules in the chlorosomal rod elements, BChl a molecules in CsmA, FMO protein, and reaction center (the double diamond represents the P840 special pair), and the primary acceptor Chl aPD in the reaction center; cubes represent [4Fe-4S] clusters in the reaction center and soluble ferredoxin; rhomboids represent [2Fe-2S] clusters in CsmI, CsmJ, CsmX, and PetC; squares represent heme groups in the reaction center, PetB, and soluble cytochrome c; double hexagons represent menaquinone molecules in the reaction center and cytoplasmic membrane; triple hexagons represent a FAD cofactor. 3. The Oxygen Evolving Complex (OEC) is assimilated by Cyanobacteria (2.7 - 2.5 Ga) The evolution of two Photosystems: The evolution of the oxygenic photosynthetic reaction center is currently under intense research scrutiny. There are two photosystems, known as Reaction Center Type I (RC1 or PSI) and Reaction Center Type II (RC II or PSII). The OEC became associated with RC II, but we find that both Photosystems are used by prokaryotes with slightly differing functions and evolutionary adaptations. Cyanobacteria use the oxygenic PSII and PSI electron transport system as does algae and plants. Anoxygenic bacteria use either RC I, or a prototype of RC II, depending on species and environment. At least one organism, Oscillatoria, has both. To understand the various distribution and functions of the two reaction centers, and how they might be related evolutionarily, John F. Allen (Dept of Biology, University of London) recently published a model to explain the appearance of the two reaction centers throughout the prokaryotes and higher plants and algae. It is now generally accepted that PSII evolved from a primitive PSI, the two reaction centers are in fact homologous in both function and structure (determined from kinetic, X-Ray and genetic studies). Dr. Allen calls his model a "redox switch hypothesis" in which the redox levels in the environment (amount of available reductants, etc) trigger a bacteria to switch from one reaction center to another. The associated light harvesting chlorophyll proteins, being integral within the bilipid membrane, are capable of serving as light harvesting complexes for both photosystems. Dr. Allen's model is presented below.
The original RC I is shown in the lower left, labeled as proto-RC. This reaction center operates with external reductants, such as hydrogen sulfide, and is capable of supplying electrons to a quinone pool, which can then recycle the energized electrons back to the reaction center while moving protons across the membrane into the periplasmic space thereby creating a protomotive force for the synthesis of ATP. In addition, the RC I provided electrons to soluble electron acceptors (ferredoxin) for use in organic synthesis. Over time and changing conditions, a second independent photosynthetic reaction center emerged, under a genetic control scheme which would favor one over the other. Allen argues that in this early stage the two reaction centers would have been cooperative, their expression genetically controlled by changing environmental conditions (alternatively, there may have been initial competition). The rise of the RC II enabled the bacteria, under conditions in which a reductant, such as H2S, was readily available to support photon-driven proton pumping through the quinone cycle into the periplasmic space for ATP synthesis. Single reaction center anaerobic phototropohs were either photolithotrophic (RC 1) or photoorganotrophic (RC II). Dr. Allen argues that metabolic flexibility may have selected those bacterial capable of providing one or both of the two types of photosystems, depending on environmental conditions. Over time bacteria appeared (such as in the present day Oscillatoria) in which both photosystems are present, and are synthesized and distributed through the membrane depending on environmental conditions encountered (Allen proposes this is carried out under redox control of gene expression). Oscillatoria limnetica has both RC I and RC II, and under conditions of low H2S, switches to RC II and performs oxygenic photosynthesis (Oscillatoria is a member of the Cyanobacteria. Note that there are other conditions, such as the heterocyst of Nostoc, in which RC II is shut off, as oxygen will interfere and shut down nitrogen fixation. See also C4 cell et al.). As the image above shows, bacteria with RC II only are known (anoxygenic), as well as bacteria with RC I only (also anoxygenic) Dr. Allen proposes that at some point in time the genetic "redox switch" was not longer needed and the two photosystem began to specialize and work in cooperation. Genetic regulatory control can still take place by either lowering or increasing either the reaction centers themselves or distributing their associated light harvesting complexes between the two. 4. The molecular structure of the manganese-calcium oxide water splitting complex [This section will be complete June 2d, in the am] Resolution: The recent 3.5 Angstrom resolution of the OEC has opened a black box that has been closed for many years. Earlier kinetic and spectroscopic data are falling in line with the mechanism of this complex but the race is on to determine exactly how the oxygen evolving complex works. We now know there are 4 manganese atoms (their respective redox states and behavior has not been determined with certainty), a calcium atom, now believed to be an essential component of the water oxidizing site, and a chloride (Cl-) co-factor. Chloride does not appear to be essential but is believed to be a charge stabilizing co-factor. An excellent animation of the Photosystem II complex, including the light harvesting chlorophylls, the reaction center and the OEC can be downloaded at Dr. Johannes Messinger's site (36 megs, Windows, RealTime) and is a beautiful rendition of PSII (highly recommended). Dr. Messinger suggests that once the mechanism of water oxidation has been determined, it may be used to engineer an energy source by combining the oxidation of water with a hydrogenase (future solar hydrogen and oxygen production). The mechanism of water oxidation has eluded scientists for decades. In the last 50 years much work has been carried out on the kinetics of PSII (OEC). In addition electrophoresis and related techniques carried out over the last 30 years had identified most of the polypeptides associated with PSII. New X-Ray diffraction studies have identified the stereochemistry and atomic structure of the OEC and surrounding complexes. It appears now that the race is on to determine the mechanism of water splitting (one of biology's greatest secrets). Once the mechanism of water oxidation is elucidated, it will become the greatest milestone of biochemistry since the discovery of DNA's structure in 1954.
From the image at left, PSII is diagrammatically depicted as a dimer, as it is found embedded in the thylakoid membranes (see Dr. Messinger's video, which clearly shows the central axis of this complex). The PSII reaction center complex is surrounded by additional light harvesting chlorophylls, shown here as LHCII (only 2 are shown, these would completely surround the complex in a natural state.
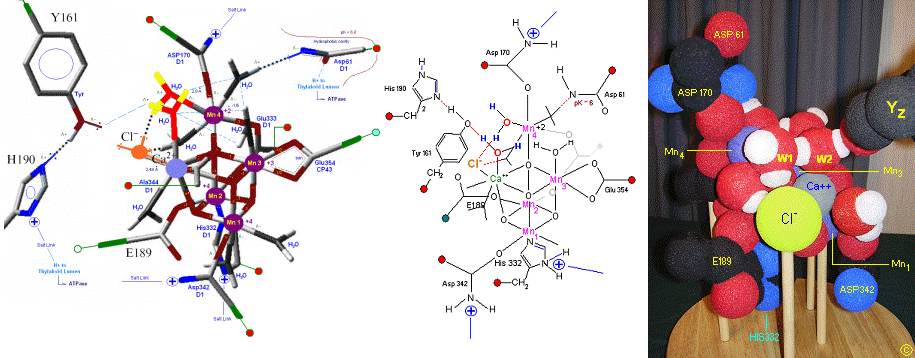
The general reaction is also outline (detailed below) in which the reaction center chlorophyll dimer, P680, is initially oxidized by an incoming photon, the electron being passed to the nearby pheophytin and then onto the quinone pool. Manganese, bound to water, calcium, chloride and oxygen (in manganese oxo bridges) then rapidly reduces Yz (a local Tyrosine) which itself became oxidized when it reduced the oxidized P680+. The basic process of water oxidation's mechanism was worked out 35 years ago by Joliot (X) and Kok (Y). Each PSII reaction center within the membrane works independently of one another and cycles through a set of 5 oxidation states, known as the "S" states (So -> S4). Our current understanding of the mechanism of water oxidation proceeds through these four states known as the "Kok Scheme" (for more details on the structure of PSII proper see the references and links provided below. The X-Ray data and model of the OEC are shown below.
The original X-Ray at a resolution of 3.5 Angstroms, was reported by Ferreira et al in 2004. The image to the left, above, was taken from an article published by James P. McEvoy, Yale University (X). The image was colored coded and the middle image drawn directly from that. The model on the right was built using this X-Ray information. The salient features of the complex (not all natural ligands may be in place, and McEvoy added an additional water ligand to complete the complex) include the manganese-calcium oxo bridging of the complex. The manganese atoms are labelled 1, 2, 3 and 4, with manganese number 4 protruding outside the main cubical structure below it, which includes manganese oxo binding, between the Mn 1, Mn 2 and Mn 3. Calcium is placed as one of the cuboidal corners (shown in blue on the left). Chloride is believed to reside on the outside of this cubane complex, and in our model on the right, appears to sit nicely (electrostatically) amongst the water molecules and calcium that surround it. Yz, the primary electron acceptor of the OEC, sits to the left, approximately 3.4 Angstroms away from the chloride. Note the numerous ligands supporting the complex, which includes several aspartates (contributed from the nearby D1 protein) as well as a histidine residue (near the bottom), also from D1. The substrate water molecules, labeled W1 and W2 in our model (right) is, at this time, an arbitrary assignment since the exact details of the mechanism is not yet known (we chose these water molecules after the model was built because of their proximity to Mn 4, and to illustrate a potential source of oxygen in our mechanistic scheme below. [when this article is complete tomorrow am (6-2) these images will be clickable to yield a closer view]. A proposed water channel may be directed out from aspartate 170 (D1) since it is believed that Tyrosine and histidine (left of the manganese cube) reside in a hydrophobic environment. [At press time this page was not complete, and will be
completed later today - final page should be up first thing on the
morning of June 2d, tomorrow (please return then for a closer look).] Water oxidizing Mechanism here: [Mechanistic proposal to be on line on morning of June 2d [page not completely finished in time for Press]
| ||
| References:
[references
not yet completely compiled] 1. Lowe, D. R., Early Life on Earth, Nobel Symposium No. 84, ed., Bengtson, S., Columbia Univ. Press, New York, 24-35 (1994) 2. Michael Russell, First Life, The American Scientist Volume: 94 Number: 1 Page: 32 (American Scientist Online) 3. Johannes Messinger, Evaluation of different mechanistic proposals for water oxidation in photosynthesis on the basis of Mn4OxCa. Physical Chemistry Chemical Physics, 2004, 6, 4764-47771. 4. Russell, M. J., Daniel, R. M., Hall, A. J., Sherringham, J. A., J. Mol. Evol., 39:231-43 (1994) 5. - 13. 14. Anbar, A. D., Holland, H. D. Geochim. Cosmochim. Acta, 56:2595-2603 (1992) 15. - 19. 19. Dawson, Scott, Creatures from the Black Lagoon: Lessons in the Diversity and Evolution of Eukaryotes, Berkeley University, 1-18 (19__) Brandes, Jay and Hazen, Bob, Key chemical in life creation - ammonia - created at Hydrothermal Vents. Carnegie Institution's Geophysical Laboratory (1998) Rasmussen, Birger, Filamentous microfossils in a 3,245-million-year-old volcanogenic massive sulphide deposit. Nature 405 676-679 (June 2000), Dept. of Geology and Geophysics, University of Western Australia, Nedlands, Western Australia 6907, Australia Filamentous microfossils in a 3,235-million-year-old volcanogenic massive sulphide deposit Blank, Biologist's Find Alters The bacteria Family Tree, Science Daily, Department of Earth & Planetary Sciences at Washington University in St. Louis Photosystem II - General references Ferreira, Kristina, Iverson, Tina, Maghlaoui, Karim, Barber, James, Architecture of the Photosynthetic Oxygen-Evolving Center, Science 303:1831-1837 (2004) Govindjee and Krogmann, David, Discoveries in oxygenic photosynthesis (1727-2003): a perspective, Photosynthesis Research 80:15-57 (2004) Haumann, Micahel, and Junge, Wolfgang, Photosynthetic Water Oxidation: A Simplex-Scheme of its Partial Reactions, Biochem. Biophys. Acta 1411:1-6 (1999)
Hillier, Warwick and Wydrzynski, Tom, The
Affinities for the Two Substrate Water Binding Sites in the O2 Evolving
Complex of Photosystem
II Vary Independently during S-State Turnover,
Biochemistry 200, 39:4399-4405 (2000)
J.P. and Shock, E.L., Energetics of overall metabolic reactions in
thermophilic and hyperthermophilic Archaea and bacteria. FEMS
Microbiology Reviews 25:175-243 (2001)
McEvoy, James P., Gascon, Jose A. et al., Computational Structural Model of the Oxygen Evolving Complex in Photosystem II: Complete Ligation by Protein, Water and Chloride, Photosynthesis: Fundamental Aspects ot Global Perspectives, A. van der Est and D. Bruce, Eds., 278-80 (2005) Messinger, Johannes, Evaluation of different mechanistic proposals for water oxidation in photosynthesis on the basis of Mn4OxCa structures for the catalytic site and spectroscopic data, Phys. Chem. Chem. Phys. 6:4764-4771 (2004) Zouni, Athina, Horst-Tobias Witt, Kern, Jan et al. Crystal structure of Photosystem II from Synechococcus elongatus at 3.8 Angstrom resolution, Nature 409:739-743 (2001)
|
 to as the "Oxygen Evolving Complex" or
OEC (also referred to as a water oxidase). The OEC is found on the oxidizing side of Photosystem II (PSII), and is
located within organelles known as
to as the "Oxygen Evolving Complex" or
OEC (also referred to as a water oxidase). The OEC is found on the oxidizing side of Photosystem II (PSII), and is
located within organelles known as  Fig 1. is an illustration of the earth relatively soon after it coalesced. Surface temperatures would have been very high, as well as meteorite impact frequency (although in time the meteor impact
frequency would decrease to about one impact per century by 4.0 Ga). (
Fig 1. is an illustration of the earth relatively soon after it coalesced. Surface temperatures would have been very high, as well as meteorite impact frequency (although in time the meteor impact
frequency would decrease to about one impact per century by 4.0 Ga). (
 Photosystem
II consists of an array of proteins, held together by numerous salt
links, hydrogen bonds, water molecules et al. and can become especially
labile under stressful conditions such as high temperatures and low
pH. The complex is also sensitive to photoinactivation if electron
flow throw the "Z" scheme is not delicately balanced (ATP
phosphorylation of the surrounding light harvesting chlorophyll proteins
is part of this regulatory scheme (
Photosystem
II consists of an array of proteins, held together by numerous salt
links, hydrogen bonds, water molecules et al. and can become especially
labile under stressful conditions such as high temperatures and low
pH. The complex is also sensitive to photoinactivation if electron
flow throw the "Z" scheme is not delicately balanced (ATP
phosphorylation of the surrounding light harvesting chlorophyll proteins
is part of this regulatory scheme (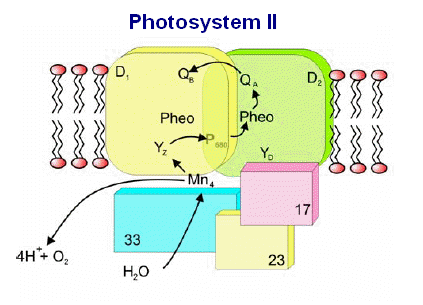 The
various proteins that surround the PSII complex are shown in the diagram
at left. Then include the principle Light Harvesting Complexes
(LHCI), D1, and D2, as well as associated peptides including the 33
kdal, 23 kdal and 17 kdal. These periphery proteins support the
reaction center and associated cytochromes and other pigments.
The
various proteins that surround the PSII complex are shown in the diagram
at left. Then include the principle Light Harvesting Complexes
(LHCI), D1, and D2, as well as associated peptides including the 33
kdal, 23 kdal and 17 kdal. These periphery proteins support the
reaction center and associated cytochromes and other pigments.