|
|
|||||||||||
World of Colour
|
The physical measurement of colour | |
Measuring Concentrations using absorbance, | |
Transition Metal Solutions, |
Rhodopsin and the eye
Basics behind Dyes and Pigments
Phosphorescence and Fluorescence
A Thermochromic example
Colour Therapy
Detection using Optical Sensors
Symbols, Glossary, Disclaimer, Credits, References, feedback..
Action 3: Book turning. This image is taken from gifs (ref. 14) and is copyright restricted according to the source given (i.e. it is not the authors' own work).
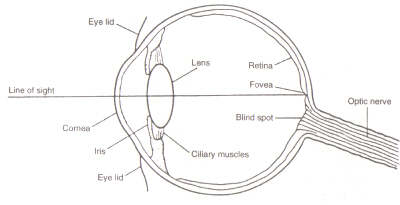
Figure 4: Structure of the eye. This image is taken from Chirstie (ref. 4) and is copyright restricted according to the source given (i.e. it is not the authors' own work).
The structure of the eye: The area within the eye (see figure 4) that detects light and colour is the called the Retina. The two types of detection cell present, rods and cones, process information coming through the Lens and send it down the optic nerve to the brain. Rod cells (of which there are around 100 million) detect the degree of lightness entering the eye and their sensitivity is dependent on the amount of Rhodopsin present which is itself generated within the cells. However, Rhodopsin is destroyed by bleaching on exposure to light and therefore rod cells only work in low light as at high illumination the reduced level of this photosensitive pigment leads to a very low sensitivity. Cone cells (of which there are around 3 million) are also sensitive to light levels but retain their function upto high illumination via use of the pigment Iodopsin. Detection of colour is a function of the three types of cone cell present within the retina: between them they cover the visible spectrum. This is because each type is sensitive to a different range of wavelengths with maximums corresponding to red (long), green (medium) or blue (short).
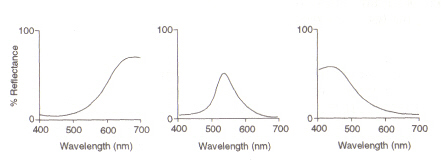
Figure 5: Maximums of (from left) red, green and blue cone cells, respectively. This image is taken from Chirstie (ref. 4) and is copyright restricted according to the source given (i.e. it is not the authors' own work).
The mechanism of Rhodopsin within the Rod cells: The protein Rhodopsin contains the protonated retinal-Schiff’s base complex which naturally lies in the inter-membrane pocket formed by the seven trans-membrane a-helical receptors. There are many flat discs of rhodopsin within the outer segment of a rod cell which upon light detection undergo a photo-isomeric change from Rhodopsin (11-cis) to all-trans retinal. After the photoisomerisation cascade which occurs via 5 shortlived intermediates (flowchart 1), trans retinal diffuses away and is converted back into 11-cis retinal before re-entry into the cycle. This process occurs via reduction to all-trans retinol followed by oxidation/isomerisation in the dark. Photoexcited Rhodopsin (4th of the 5 intermediates) triggers an enzymatic cascade process resulting in the hydrolysis of GMP. This in turn closes cation-specific channels within the rod cell membrane which are naturally open to influx of Na+ in the dark, and due to the effect of hyperpolarisation, the inner synatic body sends a nerve signal to other neurons in the Retina. Finally the light-induced lowering of calcium levels aids recovery of excited neurons to a passive, "dark" state and the cycle starts again upon detection of light. The photoreceptors of cone cells are also seven a-helical receptors with 11-cis-retinal as their chromophore. The detection range varies from green to red as the three nonpolar hydroxyl-containing residues near retinal are sequentially replaced with polar ones.

Left - Figure 6: Position of 11-cis retinal within Rhodopsin. This image is taken from Stryer (ref. 6) and is copyright restricted according to the source given (i.e. it is not the authors' own work).
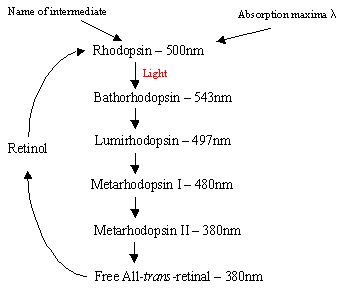
Right - Flowchart 1: The 5 intermediates of the Rhodopsin cycle.
Humans cannot make Rhodopsin, instead they use and external source, b-carotene, that is found in food in order to synthesis it:
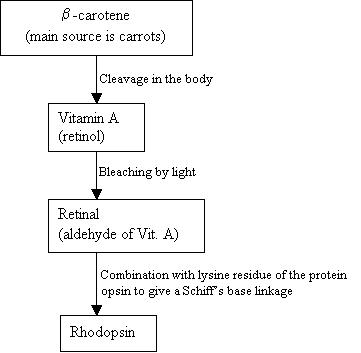
|
Flowchart 2: The synthesis of Rhodopsin New picture = Figure 7: The structures of b-carotene (top), Vitamin A (middle), and 11-cis retinal (bottom).
|
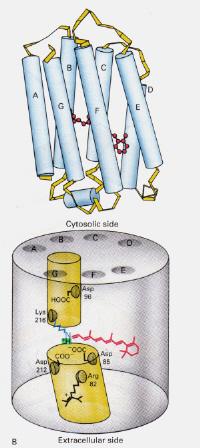
How does the mechanism of Bacteriorhodopsin found in Halobacteria differ from that of Rhodopsin found in Rod cells of the human eye?: This time the protonated retinal-Schiff’s base complex naturally blocks a channel through the membrane otherwise formed by two adjacent chambers. The protonated trans complex donates a proton to Asp-85 which then allows exit of that same proton to the extracellular side. Photoisomerisation to the 13-cis structure allows the Schiff’s base to pick up a proton from the Asp-96 residue on the cytosolic side. Upon reorientation of the cis form to the trans, the cycle of isomerisation and proton pumping continues.
Figure 8: The structure of Bacteriodopsin. This image is taken from Stryer (ref. 6) and is copyright restricted according to the source given (i.e. it is not the authors' own work).